Struggling with Unreliable Angiotensin Data?
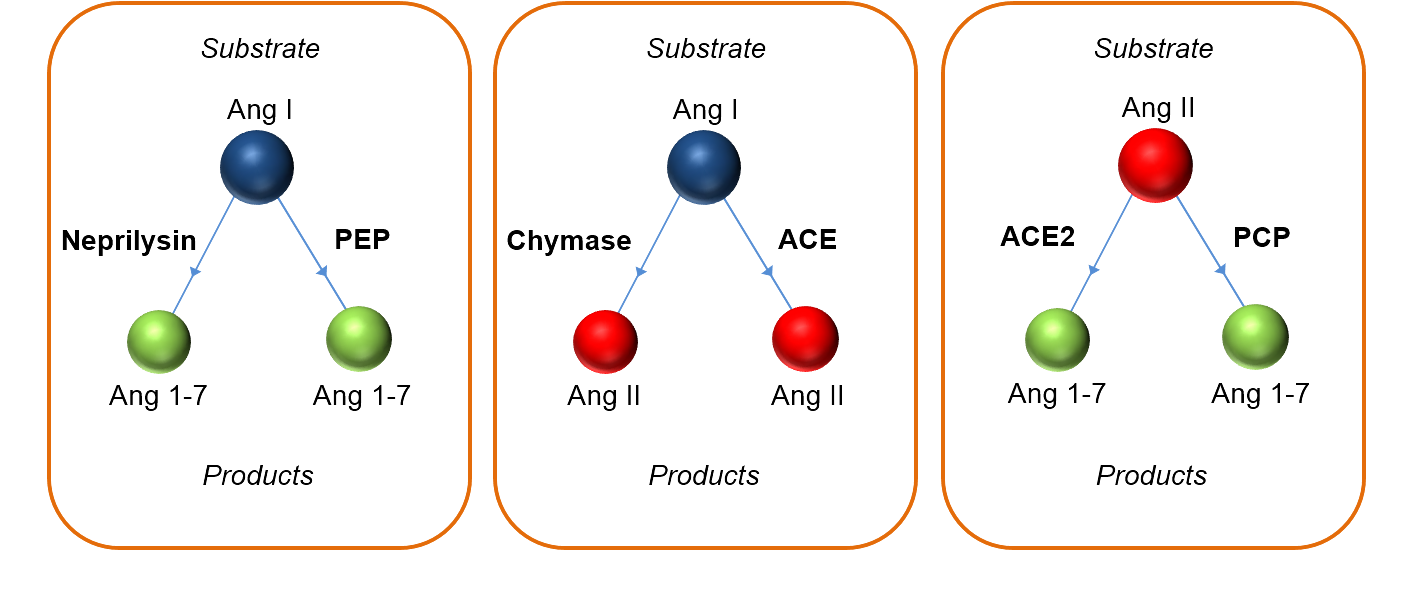
Short half-lives, ongoing formation, and cross-reactive antibodies make angiotensin measurement notoriously tricky — and often inconsistent across studies.
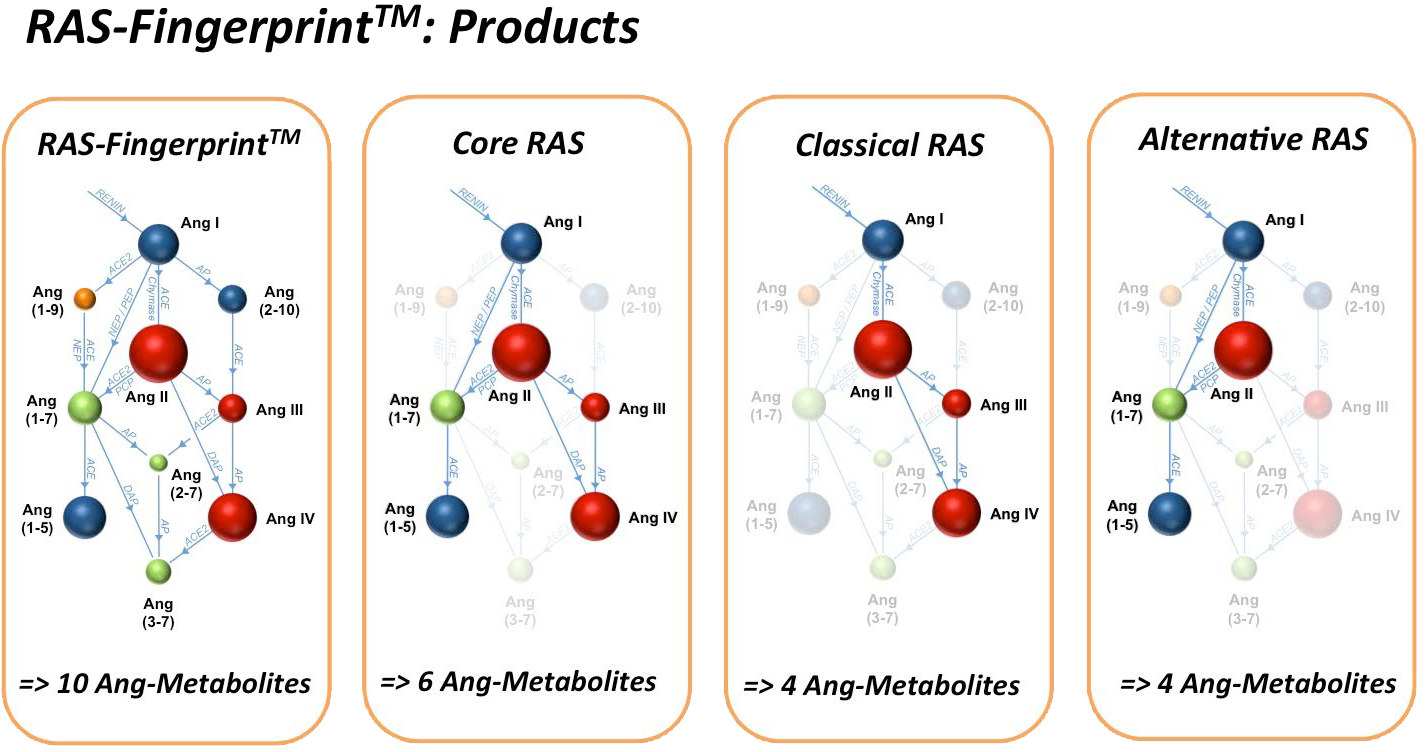
Meet the Game-Changer: RAS-Fingerprint™
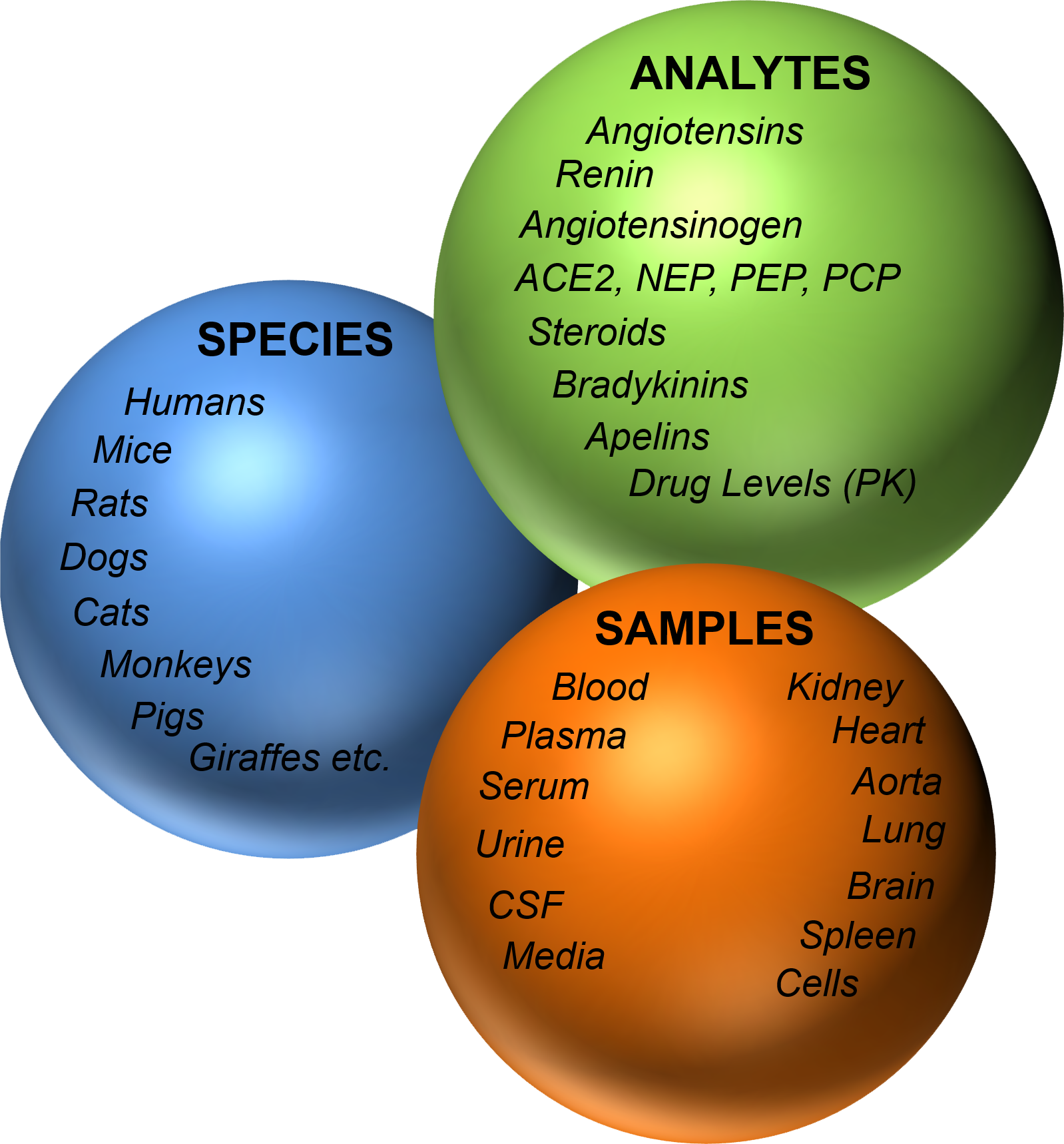
Our RAS-Fingerprint™ service solves these issues at the root. With expert-guided sampling, optimized inhibitors, and gold-standard LC-MS/MS analysis, we analyze plasma or tissue to deliver precise, reproducible data — plus clear, eye-catching visualizations that make your results impossible to ignore.